The final RNAiL?
A recent article in The Scientist asks whether, in light of a paper by Lin et al. showing phenotypic discrepancies between RNAi and CRISPR, this is not ‘the last nail in the coffin for RNAi as a screening tool’?
The paper in question found that a gene (MELK) that had been shown by many RNAi-based studies to be critical for several cancer types shows no effect when knocked out via CRISPR. They also report that in relevant published genome-wide screens, MELK was not at the top of the hit lists.
Does this mean that the papers that used RNAi were unlucky and off-target effects were responsible for their observed phenotypes?
Gray et al. identified MELK as a gene of interest based on microarray experiments. They then designed RNAi experiments to test its role in proliferation. Assuming that this study and the subsequent ones followed good RNAi experimental design (using reagents with varying seed sequences, testing the correlation between gene knockdown and phenotypic strength, etc.), we can be fairly confident that MELK is involved in proliferation. It might not be the most essential player, which would explain why it is not at the top of screening hit lists. And screening lists have the draw-back of enriching for off-target hits.
Another possibility is that Lin et al. have observed a known complicating feature of knock-out screens: genetic compensation. Although they undertake experiments to address this issue, it could be that compensation takes place too quickly for their experiments to rule it out. Furthermore, they could have addressed this issue by testing knock-down reagents themselves, and checking whether genes they hypothesise as responsible for the supposed off-target effect in the published RNAi work are in fact down-regulated. C911 reagents could also be used to test for off-target effects. This is extra work, but given that they are disputing the results in many published studies, this seems justified.
As regards the role of RNAi in screening, The Scientist concludes with the following (suggesting that their answer to the question of whether this is the final nail is also No):
In the meantime, one obvious solution to the problem of target identification and validation is to use both CRISPR and RNAi to validate a target before it moves into clinical research, rather than relying on a single method. “We have CRISPR and short hairpin reagents for every gene in the human genome,” said Bernards. “So when we see a phenotype with CRISPR, we validate with short hairpin, and the other way around. I think that would be ideal.”
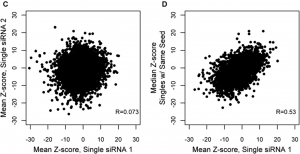
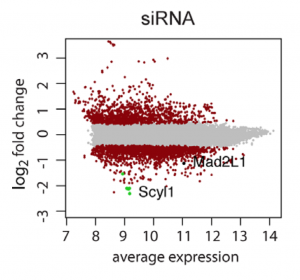
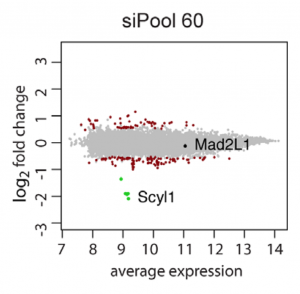
Although we agree that validating CRISPR hits with RNAi reagents is important (especially if drugability is a concern), one has to be careful with RNAi reagents, like single siRNAs/shRNAs or low-complexity pools, that are susceptible to seed-based off-target effects. For validating CRISPR screening hits, siPOOLs provide the best protection against unwanted off-target effects, saving you time, money, and disappointment during the validation phase.
Want to receive regular blog updates? Sign up for our siTOOLs Newsletter: