Seed effects persist in hyperdimensional space
Work from the Carpenter lab suggests that attempts to shake seed-based off-targets by going to ‘phenotypic hyperspace’ will not work.
They performed a high-content assay with 315 shRNAs covering 41 genes. A 1301-dimensional profile was created for each well, and compressed to 205 principal components that captured 99% of the variance.
The hope would be that by examining a wider phenotypic space, the gene-specific effects of RNAi reagents would become more prominent.
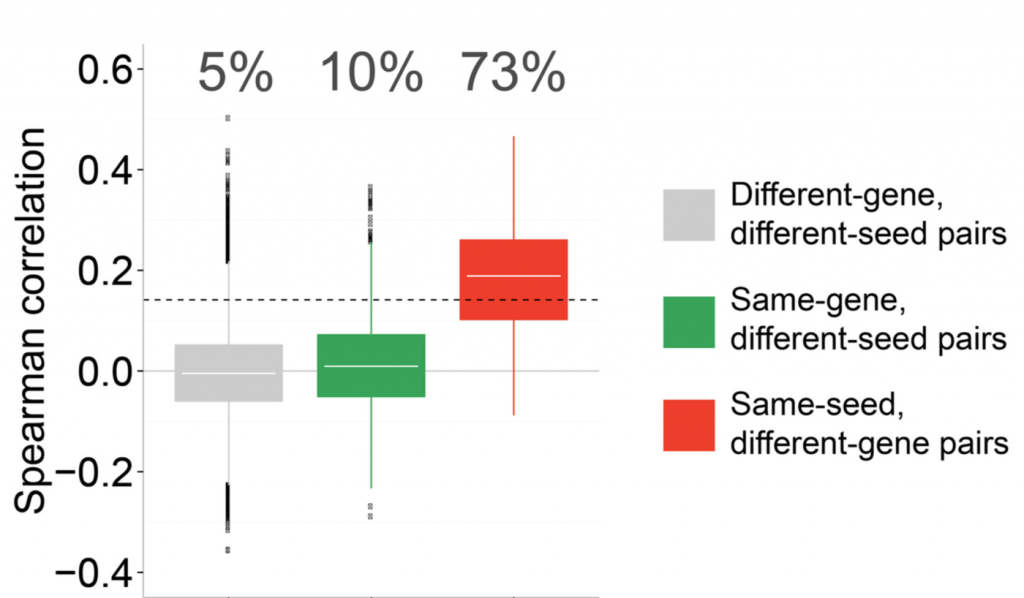
However, the profiles between shRNAs targeting the same gene are only slightly better than those between random shRNAs, while shRNAs sharing the same seed sequence have much more similar profiles.
(figure shows percent of significant profile correlations for different pairings)
Off-target phenotypes can only be escaped by using a reagent that exclusively knocks down the target gene.
Want to receive regular blog updates? Sign up for our siTOOLs Newsletter: