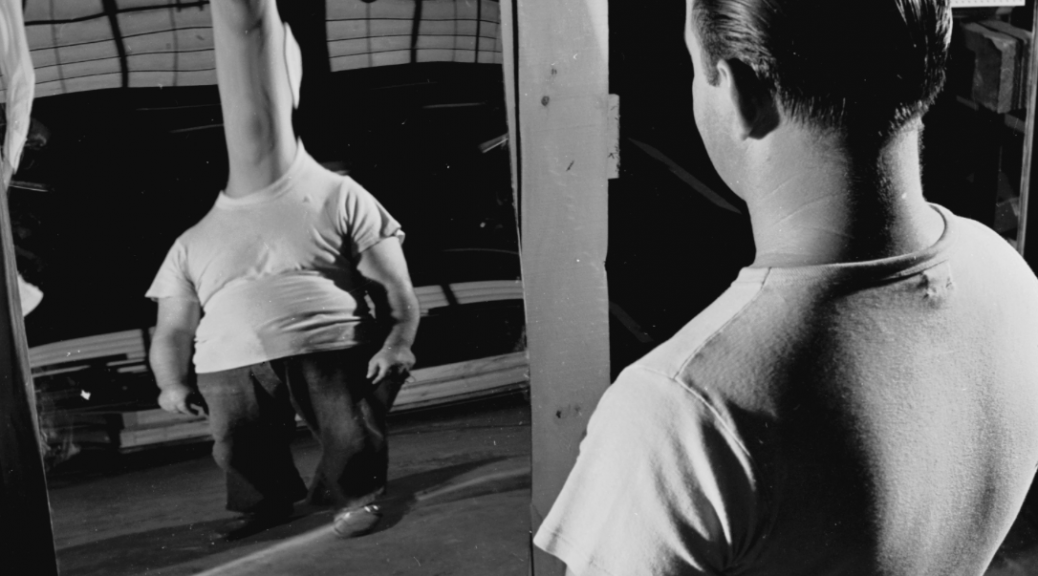

Little correlation between Dharmacon siGENOME and ON-TARGETplus reagents
The most common way to validate hits from Dharmacon siGENOME screens is to test the individual siRNAs from candidate pool hits (siGENOME reagents are low-complexity pools of 4 siRNAs). In this deconvolution round, we normally see that the individual siRNAs for genes behave very differently and seed effects dominate (discussed here and here).
One could argue that deconvolution is not the correct way to validate candidate hits (even though it’s the method recommended by Dharmacon), as testing the siRNAs individually will result in seed effects that are suppressed when the siRNAs are pooled. One problem with this argument is that low-complexity pooling does not get rid of off-target effects (e.g. Fig 5 in this paper), something that is better done via high-complexity pooling. But assuming it were true, validating with a second Dharmacon pool would be better.
Tejedor et al. (2015) performed a genome-wide Dharmacon siGENOME screen for regulators of Fas/CD95 alternative splicing. ~1500 genes were identified by a deep-sequencing approach. ~400 of those were confirmed by high-throughput capillary electrophoresis (HTCE, LabChip). They then retested those ~400 genes (again by HTCE) using Dharmacon ON-TARGETplus pools.
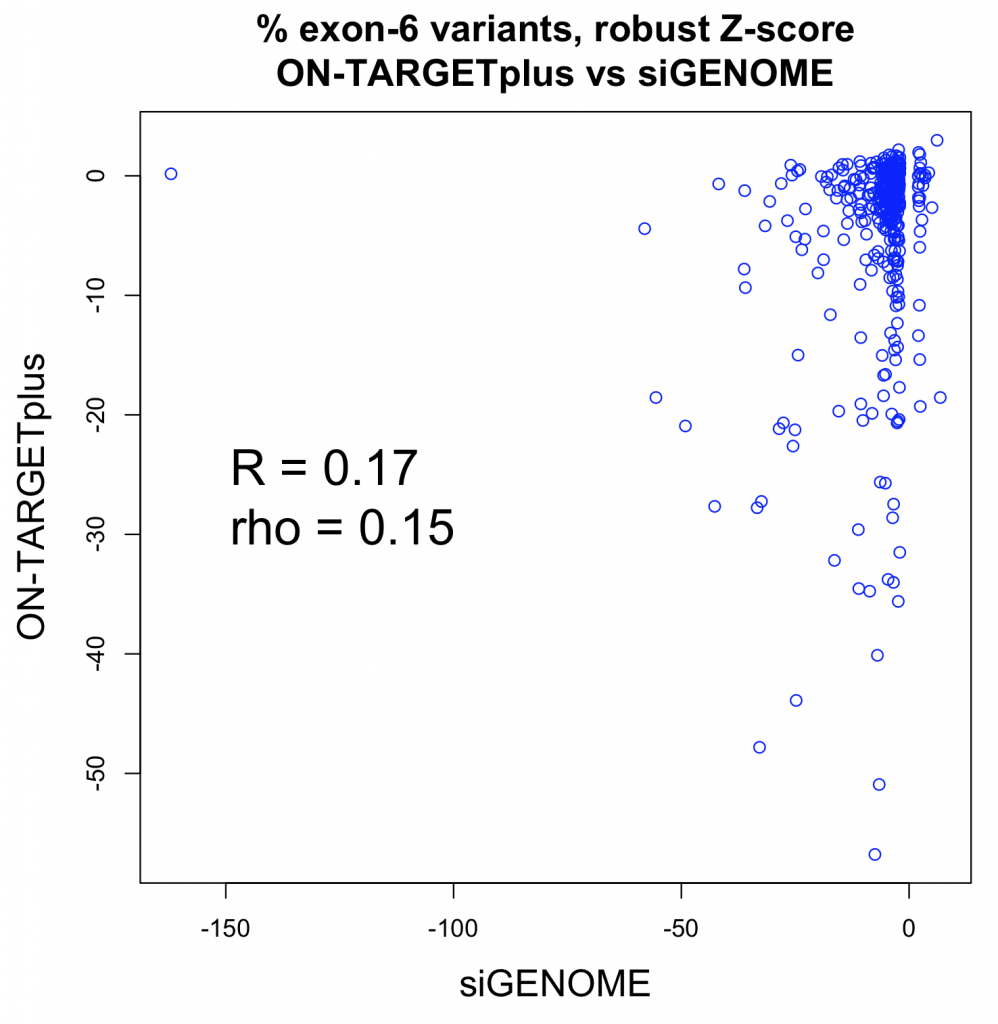
The following plot shows the values for the siGENOME and ON-TARGETplus pools for the same genes (i.e. each point corresponds to 1 gene).
What’s measured is the percent of splice variants that include exon 6 following siRNA treatment. That was compared to the values for a plate negative control (untransfected wells) and converted to a robust Z-score. This is the main readout from the paper.
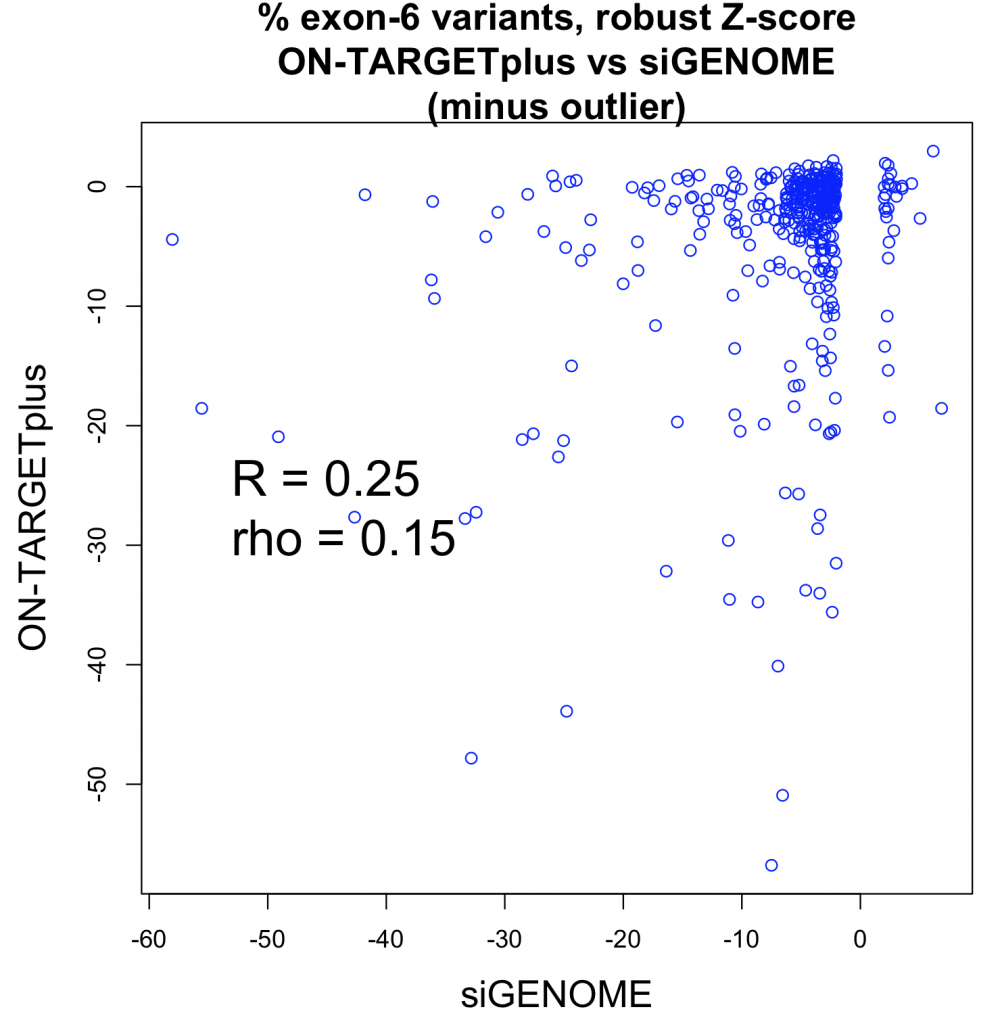
The Pearson correlation improves if the strong outlier at -150 for siGENOME is removed (R = 0.25), while the Spearman correlation is unchanged.
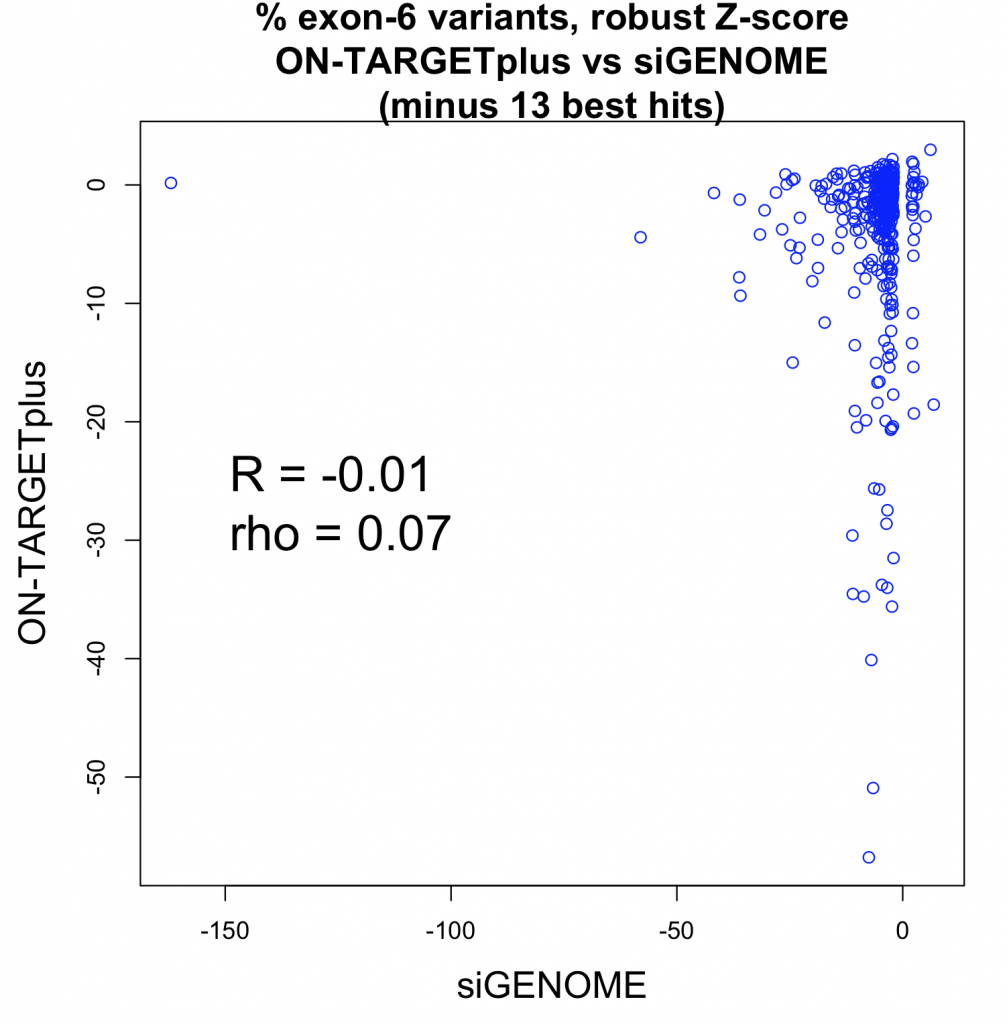
We see that a fairly small number of genes are giving reproducibly strong phenotypes (e.g. 13 of 400 have robust Z-scores less than -15 for both siGENOME and ON-TARGETplus reagents).
If we remove those 13 strong hit genes, the correlation approaches zero:
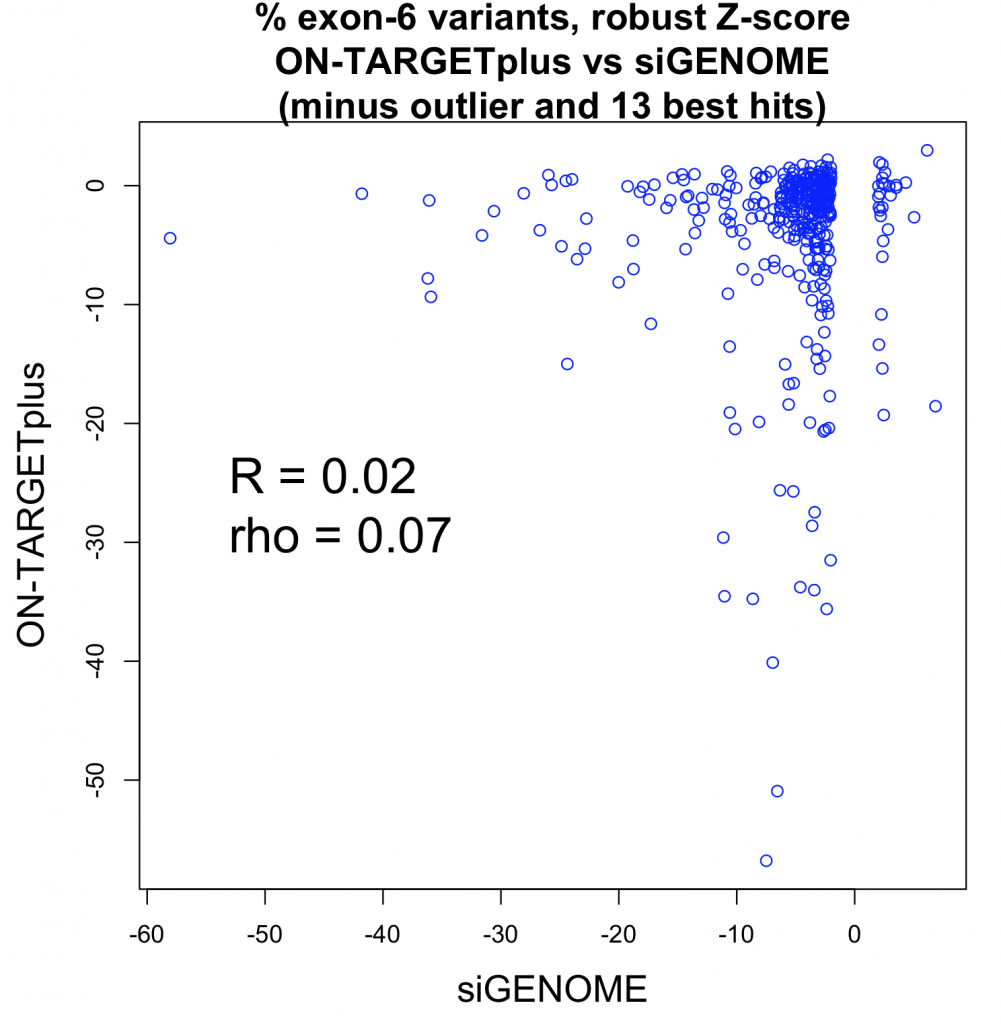
Even if the strong outlier for siGENOME is removed, the correlation is still near zero:
Although using a second Dharmacon pool removes some of the arbitrariness of defining validated hits (e.g. saying that 3 of 4 siRNAs must exceed a Z-score cut-off of X, or 2 of 4 siRNAs must exceed a Z-score cut-off of Y), the end result is similar: A few strong genes show reproducible phenotypes, while many of the strongest screening hits show inconsistent results. The main problem, off-target effects in the main screen, is not fixed.
postscript
Tejedor et al. say that 200 genes were confirmed by ON-TARGETplus validation. They consider a gene confirmed if the absolute value of the robust Z-score is greater than 2. The Z-score is calculated using the median for untransfected plate controls. I suspect that a significant proportion of randomly selected genes would also have passed this cut-off.
In table S3 (which has the ON-TARGETplus validation results), there are actually only 177 genes (including 2 controls) that meet this cutoff. The supplementary methods state: Genes for which Z was >2 or <-2 were considered as positive, and a total number of 200 genes were finally selected as high confidence hits.
Which suggests that genes outside the cut-off were chosen to bring the number up to 200.
But if we look at the Excel sheet with the ‘200 hit genes’, it has 200 rows, but only 199 genes. The header was included in the count.
This type of off-by-one error is probably not that uncommon. In a case like this, it does not matter so much.
One case where it did matter was in the Duke/Potti scandal. The forensic bioinformatics work of the heroes of the Duke scandal found that, when trying to reproduce the results from published software, one of the input files caused problems because of an off-by-one error created by a column header. That was one of many difficulties in reproducing the Potti paper’s results which eventually led to its exposure.





2 thoughts on “Little correlation between Dharmacon siGENOME and ON-TARGETplus reagents”