
“Phenoville” – RNAi & CRISPR Screening Strategies
Pleasantville is a movie based on an interesting idea: two teenagers are magically transported through their TV to a town called Pleasantville set in the 1950s where everything is perfect (and also black-and-white). As they discover the complex, imperfect emotions hidden below the idyllic surface, the black-and-white characters and objects start to gain colour.
In loss-of-function genetic screening, some reagents and screening formats may also give rise to a narrow, black-and-white view of a biological process. A sort of “Phenoville”. This was illustrated nicely in a recent review of screening strategies for human-virus interactions by Perreira et al. (2016).
The authors performed screens for human rhinovirus (HRV) infection using arrayed RNAi reagents (siRNAs) and pooled CRISPR reagents (sgRNAs), and then compared the resulting hit lists.
The arrayed RNAi screen produced over 160 high-confidence candidate genes, whereas the CRISPR screen only found 2. The authors comment:
“The comparison of these two screening approaches side-by-side, using the same cells and virus, raises an interesting point. The number of host factors found for HRV14 was far greater using the MORR/RIGER approach [i.e. RNAi performed with multiple orthologous RNAi reagents and analysed by RNAi gene enrichment ranking method] and is approaching a systems level understanding based on bioinformatic analyses and the near saturation of, or enrichment for, multiple complexes and pathways (Fig. 4) (Perreira et al., 2015). By comparison our matched pooled CRISPR/Cas9 screen for HRV-HFs yielded two high-confidence candidates based on reagent redundancy, ICAM1, the known receptor for HRV14, and EXOC4, a gene involved in exocyst targeting and vesicular transport (He & Guo, 2009). Given the known role of ICAM1 as the host receptor for most HRVs, these results point to entry as the major viral lifecycle stage interrogated by a pooled functional genomic screening approach using a population of randomly biallelic null cells infected by a cytopathic virus.”
In simple terms, RNAi screening produced a richer data set that revealed system level interactions whereas CRISPR screening yielded a small number of specific hits that only affected an early-stage pathway. The ‘systems level understanding’ is nicely shown in the following diagram of the RNAi hits. The red box at the top left is the only gene (ICAM1) that was common to the RNAi and CRISPR screens.
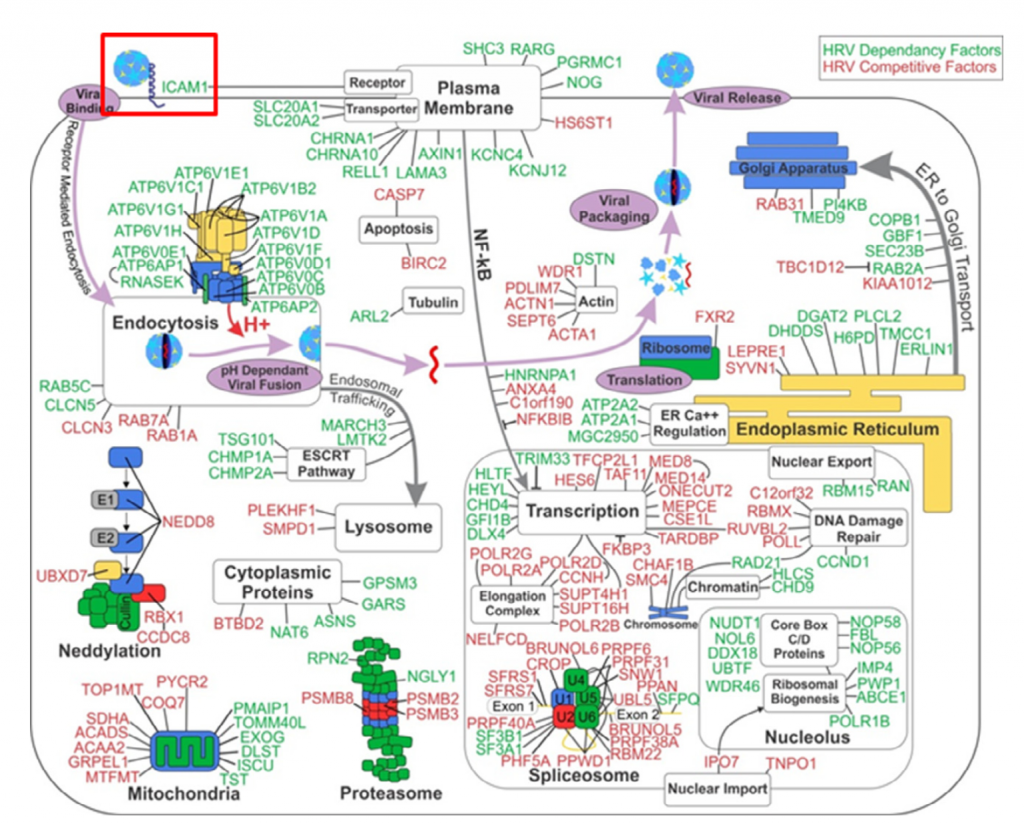
Perreira et al. conclude that arrayed siRNA screens permit the detection of a larger number of viral dependency factors, albeit with a significant tradeoff in a greater number of false positive hits (mainly due to off-target effects). In contrast, pooled screens with CRISPR sgRNAs using cell survival as a readout, as also seen with most haploid cell screens, display limited sensitivity but excellent specificity in finding host genes that act early on in viral replication (e.g. ICAM1).
In Perreira et al.‘s words:
“… given the currently available functional genomic strategies if the goal is to find viral entry factors (e.g., host receptors) with high specificity its best to use a pooled survival screen, but alternatively if the aim is to obtain with relative ease a more comprehensive set of host factors, albeit with more prevalent false positives, than an arrayed siRNA screen would be the preferred method.”
Summarizing two options for genetic screeners:
- Arrayed RNAi screens
- provide a richer view of the underlying biology
- produce more false positives from OTEs
- produce false negatives from OTEs
- Pooled CRISPR screens
- provide a narrower view of the underlying biology
- produce fewer false positives
- produce false negatives because of genetic compensation
Off-target effects (OTEs) are the primary cause of false positives, and the resultant higher assay noise also increases the number of false negatives in arrayed RNAi screens. Reagents like siPOOLs minimize the risk of off-target effects and reduce assay noise.
One key factor not mentioned by Perreira et al. is the presence of genetic compensation in gene knockout approaches.
Putting genetic compensation in terms of human actors, imagine that you are investigating the function of bus drivers in Pleasantville. To induce loss-of-function, assume that aliens will be abducting the bus drivers. If the bus drivers are abducted in their sleep (equivalent to a CRISPR knock-out), you may not get a good idea of their function when you film the next day. People may be compensating by driving, biking or staying home. Alternatively, the bus company may have found emergency replacement drivers.
Now suppose the bus drivers are abducted in the middle of the day while driving their routes (equivalent to an RNAi knock-down). The film will show buses crashing (hopefully without any serious injuries, since this is just a TV show!) and the public transportation system will suddenly come to a halt.
RNAi gene knockdown screens with siPOOLs can provide a significant advantage over CRISPR gene knockout screens in obtaining a system level understanding in biological models.
Want to receive regular blog updates? Sign up for our siTOOLs Newsletter: