Is it important to avoid microRNA binding sites during siRNA design?
Summary: To address the question of whether one should avoid microRNA binding sites during siRNA design, we examined whether removing siRNAs that share seeds with native microRNAs would reduce the dominance of seed-based off-target effects in RNAi screening.
siRNA design and native microRNA target sites
Recently, we discussed a review of genomics screening strategies. The authors state:
RNAi screens are powerful and readily implemented discovery tools but suffer from shortcomings arising from their high levels of false negatives and false positives (OTEs) as can be seen when comparing the low concordance among the candidate genes detected in different screens using the same species of virus, e.g., HIV-1, HRV, or IAV (Booker et al., 2011; Bushman et al., 2009; Hao et al., 2013; Perreira et al., 2015; Zhu et al., 2014).
To address these concerns, improvements in the design and synthesis of next-gen RNAi library reagents have been implemented including the elimination of siRNAs with seed sequences that are complementary to microRNA binding sites.
Given that off-target effects via microRNA-like binding are the main source of RNAi screening phenotypes, avoiding native microRNA sites during siRNA design seems like a reasonable strategy. But does it make much difference in actual RNAi screens?
Hasson et al. 2013 performed a mitophagy screen using the Silencer Select siRNA library. About 12% of the ~65,000 screened siRNAs have a 7-mer seed shared by a miRBase microRNA.
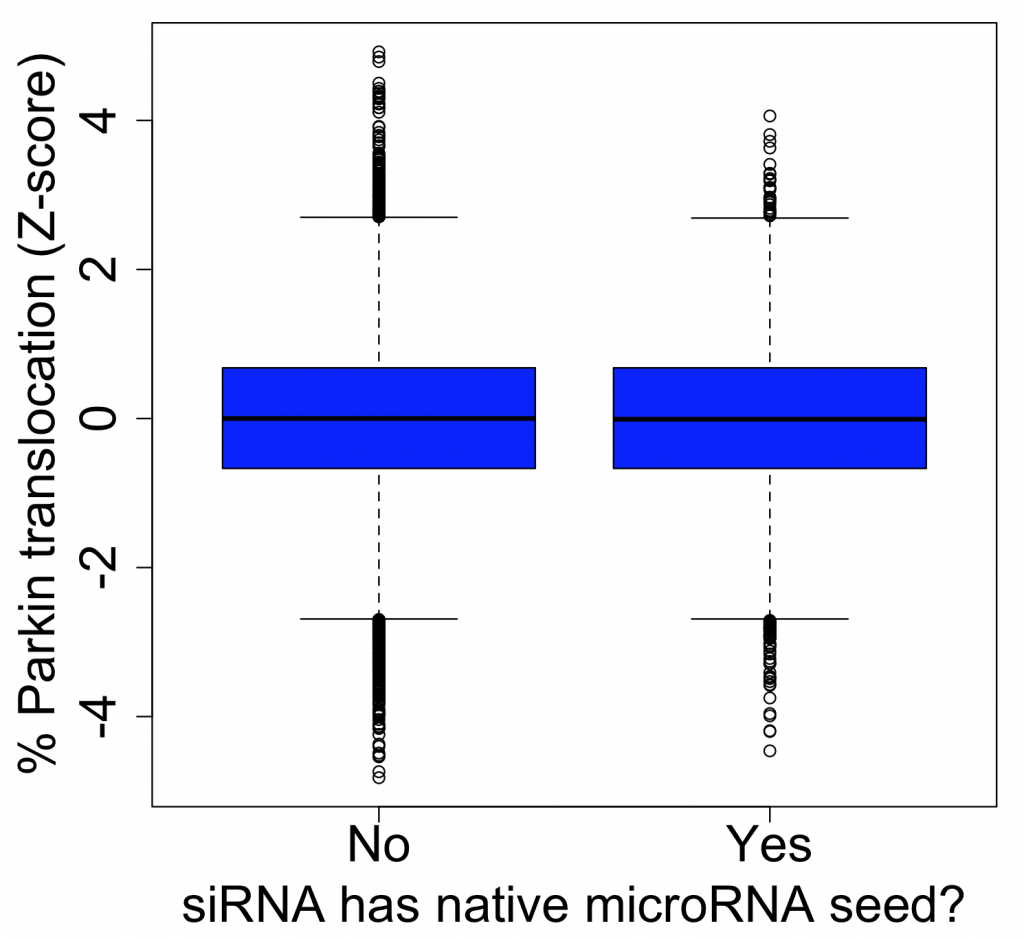
The screen’s main phenotypic readout, % Parkin translocation (PPT), is strongly affected by seed effects. The intra-class correlation for siRNAs with the same seed is ~.51 (versus ~.06 for siRNAs with the same target gene). There appears to be no difference between how siRNAs with or without microRNA seeds behave:
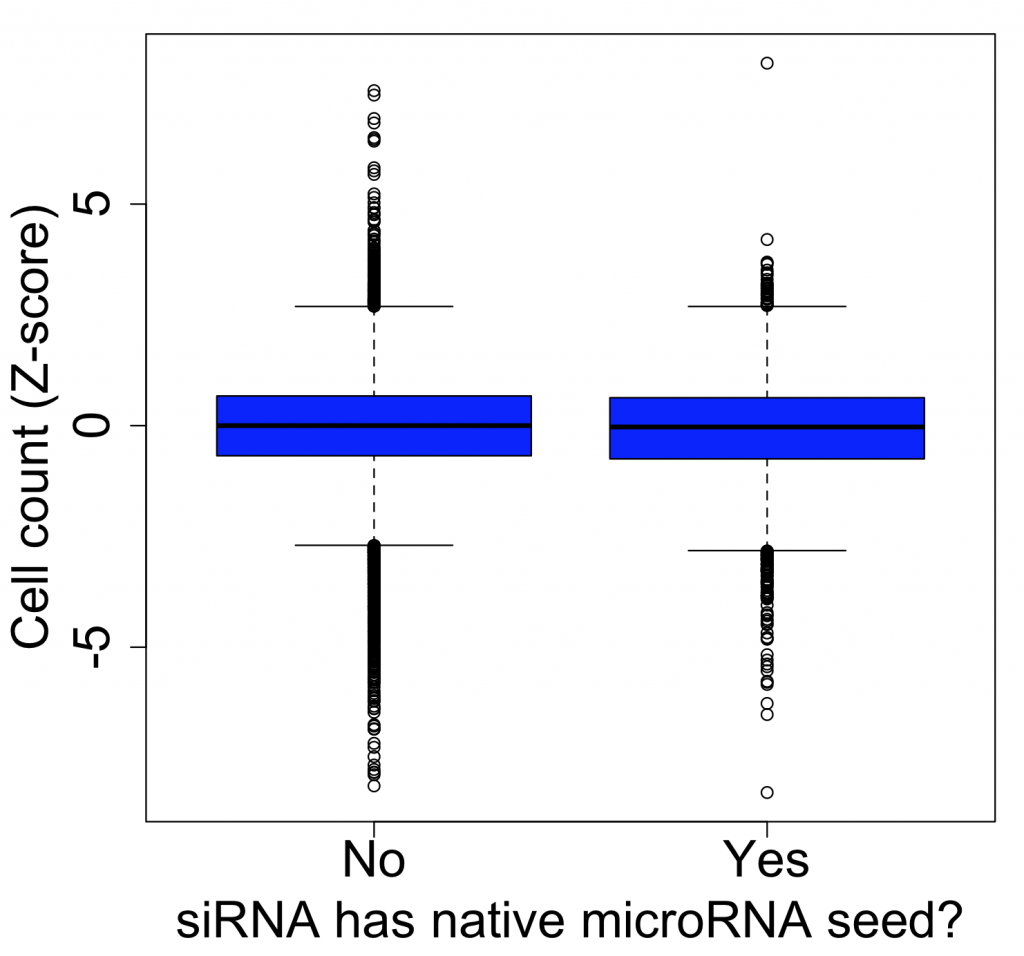
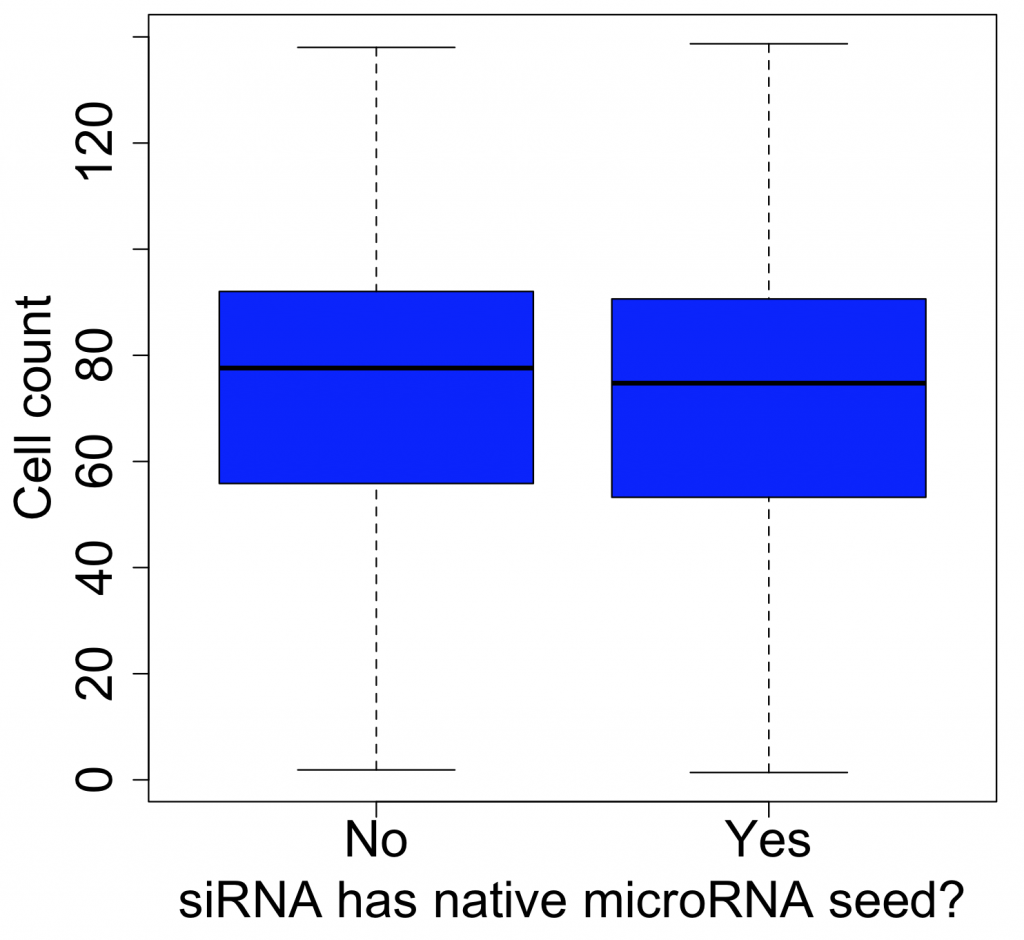
The same thing is found if we look at a less specific phenotype like cell count (which should be more broadly susceptible to off-target effects, as more genes should affect this phenotype):
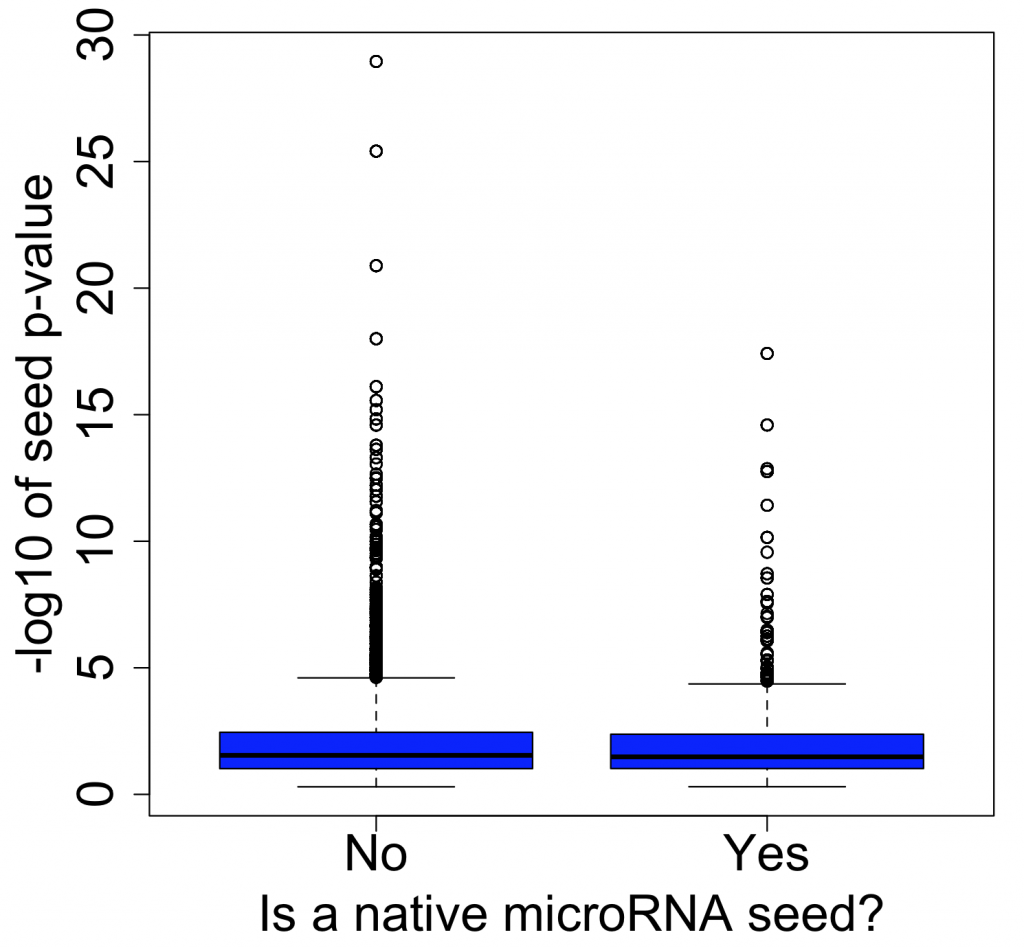
And if we look at seeds that are enriched at the top of the screening list (sorted by descending PPT), we also don’t see much difference between siRNAs with or without native microRNA seeds. (Note that the seed p-value is calculated in a similar way to RSA, based on how over-represented a seed is towards the top of a ranked list)
We also examined a general phenotypic readout (cell viability) in a dozen large-scale RNAi screens.
For some screens, we do see a slight shift in the values for siRNAs with or without native microRNA seeds.
For example, a genome-wide screen of Panda et al. 2017 (also using the Silencer Select library) shows a slight decrease in viability for siRNAs with native microRNA seeds:
Removing those siRNAs does not change the dominance of seed-base off-targets.
The intra-class correlation (ICC) for siRNAs with the same 7-mer seed is ~.53, with or without the inclusion of siRNAs with native microRNA seeds, while ICC for siRNAs with the same target gene is only ~.06.
Coming back to the quote from the review article on genomic screening, next-gen RNAi library reagents that avoid native microRNA seeds are not expected to be much better than siRNAs that include them.
The most effective way to avoid seed-based off-target effects is to use high-complexity siRNA pools (siPOOLs). Learn more about siPOOLs