CRISPR/Cas9 Screening – The “Copy-Number Effect”
Several CRISPR/Cas9 screens identifying essential genes in cancer cell lines have been performed to date (Shalem et al., 2014, Hart et al., 2015, Kiessling et al., 2016). These typically take the form of pooled screens where sgRNA libraries targeting all genes or subsets of genes are introduced in parallel into Cas9-expressing cells, at a single sgRNA per cell. The sgRNAs exert a negative or positive selection pressure on cells based on their impact on cell viability and proliferation. The most depleted or enriched sgRNA sequences are determined by next-generation sequencing, revealing relevant gene ‘hits’. Very similar to how pooled shRNA screens are performed.
From these screens, several groups have observed a worrying phenomenon: CRISPR gRNAs targeting genomic regions of high copy number amplification showed a striking reduction in cell proliferation/survival. Dr William Hahn’s group at the Dana Farber Institute was one of the first to characterize this in a publication last year involving a CRISPR/Cas9 screen on 33 cancer cell lines looking for essential genes. In total, 123411 unique sgRNAs were used targeting 19050 genes (6 sgRNAs/gene), 1864 miRNAs and 1000 non-targeting negative control sgRNAs.
What they discovered is a little worrying to say the least.
The figure shows two genomic regions in two different cell lines (SU86.86 and HT29). At genomic coordinates highlighted by the red box, 3 tracks are shown. Top, copy number from the Cancer Cell Line Encyclopaedia (CCLE) SNP arrays, red indicating above average ploidy and blue showing below; middle, CRISPR/Cas9 guide scores with purple trend line indicating the mean CRISPR guide score for each CN segment defined from the above track; bottom, RNAi gene-dependency scores. AKT2 and MYC, known driver oncogenes at these loci, respectively, are highlighted in orange. For RNAi data, shRNAs targeting AKT2 used in Project Achilles were not effective in suppressing AKT2 (hence the negative result).
Key findings:
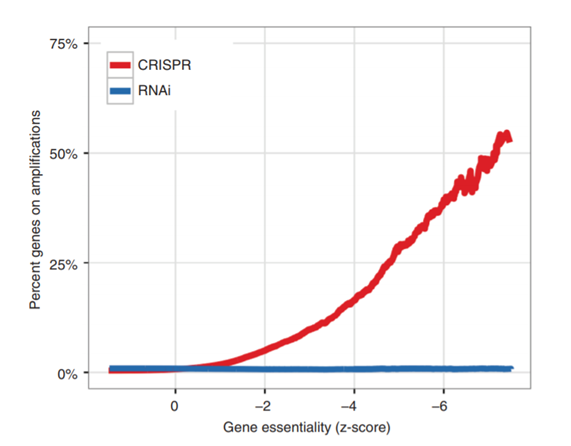
- A striking enrichment of negative CRISPR guide scores (i.e. sgRNAs that reduced cell proliferation/survival) for genes that reside in genomic regions of high copy-number amplification.
- Genes identified in CRISPR that reduced survival, did not have the same effect when disrupted by RNAi in the same cell lines (this RNAi screen was done by the same group but published 2 years before).
- This enrichment was seen also for unexpressed genes, i.e. genes not transcribed. Meaning the reduced survival was not due to loss-of-function of the targeted gene.
- Even for regions with low absolute copy numbers, a significant reduction in survival was observed compared to non-targeting control sgRNAs. Furthermore, the effect was dose-dependent with greater copy number amplifications producing larger negative CRISPR guide scores.
Notably, the correlation between copy number and genes that were scored high on essentiality was also observed when looking at data from other studies (Hart et al., 2015). The “copy number effect” would therefore produce a high number of false positives in CRISPR screens for essential genes in cancer cell lines. The graph above shows just how big an effect this is. Comparing genes identified as essential in a CRISPR screen vs RNAi screen, increasingly essential CRISPR-identified genes were more likely to reside on copy number amplifications (defined as having average sample ploidy > 2). This effect was notably absent for RNAi-derived essential genes.
Aside from false positives, the increased noise due to “copy number effects” also increases false negatives. MET, a gene identified by shRNA screens, for example, failed to be picked out by CRISPR screens as it is located on a chromosome 7 amplicon (7q31) in MKN45 cells (gastric cancer cell line) where all other gRNAs within that amplicon also scored as essential.
The authors go on to explore mechanisms behind the “copy number effect”. They found it was attributed to a DNA damage response stimulated by excessive cutting by Cas9. This response appeared p53-dependent and induced cell cycle arrest at the G2 phase, explaining the anti-proliferative effect. A similar response was seen for promiscuous sgRNAs that cut at multiple sites, with effects being more pronounced when cuts were spread over several chromosomes as opposed to a single chromosome.
How to manage this?
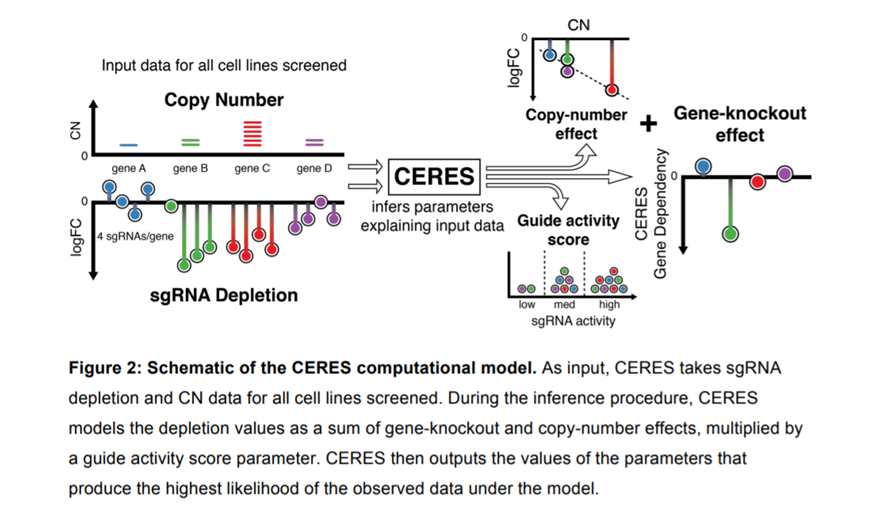
So far, most simply avoid analysing hits where sgRNAs lie at amplified regions or target multiple sites (Wang et al., 2017). However, these regions of copy number amplifications have been implicated in cancer and may contain relevant hits. Several computational methods have therefore recently been developed to correct for “the copy number effect”. Hahn’s group developed a computational algorithm called CERES based on data obtained from CRISPR sgRNA screens in 342 cancer cell lines representing 27 cell lineages.
Novartis also developed a Local Drop Out (LDO) algorithm that corrects obtained data based on examining gRNAs scores at direct genomic neighbours. When multiple neighbouring genes show similar drop out scores, effects are assumed to be due to “copy number effects”. This method has the advantage of not requiring prior knowledge of copy number, however it does require a sufficient density of gRNAs to accurately capture “copy number effects”. They also had an alternative method, Generalized Additive Model (GAM) where copy number was taken into account.
How the CERES Model Works
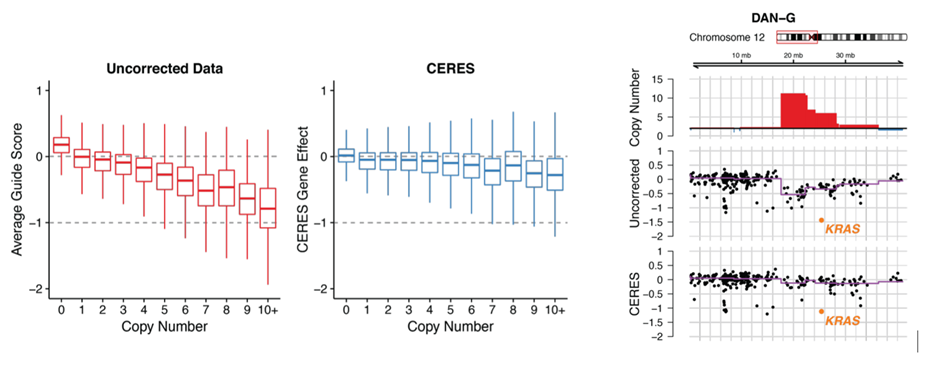
The Results – copy number dependency is reduced while preserving essentiality of cancer-specific genes such as KRAS
A step towards the right direction but the penetrance of this effect still raises some concerns:
- Although false positives are reduced with these computational methods, it is difficult to recapture false negatives. This is dependent on the gRNA having a stronger phenotype compared to neighbouring gRNAs on the amplicon which is not always the case. The LDO method for example still failed to recapture MET.
- Guide scores can vary with cell line, sgRNA and experimental conditions, making it difficult to apply the same counter-measures to every experiment.
- Given multiple cut sites trigger the same effect, how do we ensure multiple sgRNAs when introduced into a cell are not inducing a similar response? This is difficult to control in pooled screens, and poses a limitation in multiplex screens. Synthetic lethality screens for example with sgRNAs targeting multiple genes, might be subject to a higher false positive rate.
- With even diploid genes (copy number = 2) having statistically significant growth reduction compared to haploid gene loci, the challenge still remains to delineate a true loss-of-function over a non-specific cellular response.
- Negative sgRNA controls have to be carefully selected. From the study, non-targeting controls had little impact on viability compared to most other sgRNAs. Controls targeting non-expressed genes or non-essential loci have been recommended as better controls.
- Lastly, although this effect seems to apply mostly to cancer cell lines that undergo a high rate of gene amplifications, similar effects may extend to polyploid tissues such as the liver.
Hence as always gene function should be determined by a variety of methods. Using RNAi for example to affirm a CRISPR-knockout phenotype would add greater confidence to a hit. To avoid those RNAi-related false positives however, its probably best to use siPOOLs.
Source of figures:
Aguirre, A. J., Meyers, R. M., Weir, B. A., Vazquez, F., Zhang, C.-Z., Ben-David, U., … Hahn, W. C. (2016). Genomic Copy Number Dictates a Gene-Independent Cell Response to CRISPR/Cas9 Targeting. Cancer Discovery, 6(8), 914 LP-929.
Meyers, R. M., Bryan, J. G., McFarland, J. M., Weir, B. A., Sizemore, A. E., Xu, H., … Tsherniak, A. (2017). Computational correction of copy-number effect improves specificity of CRISPR-Cas9 essentiality screens in cancer cells. bioRxiv. Retrieved from https://biorxiv.org/content/early/2017/07/10/160861.abstract
Other relevant sources:
Munoz, D. M., Cassiani, P. J., Li, L., Billy, E., Korn, J. M., Jones, M. D., … Schlabach, M. R. (2016). CRISPR Screens Provide a Comprehensive Assessment of Cancer Vulnerabilities but Generate False-Positive Hits for Highly Amplified Genomic Regions. Cancer Discovery, 6(8), 900 LP-913. Retrieved from https://cancerdiscovery.aacrjournals.org/content/6/8/900.abstract
de Weck, A., Golji, J., Jones, M. D., Korn, J. M., Billy, E., McDonald, E. R., … Kauffmann, A. (2017). Correction of copy number induced false positives in CRISPR screens. bioRxiv. Retrieved from https://biorxiv.org/content/early/2017/06/23/151985.abstract
Want to receive regular blog updates? Sign up for our siTOOLs Newsletter: