
Deconvoluted SMARTpools are like a box of chocolates
Falkenberg et al. (2014) performed a synthetic lethal RNAi screen to identify genes which, when knocked down in combination with drug treatment, induced apoptosis in drug-resistant cells.
The first screening pass covered over 18,000 protein-coding genes using Dharmacon’s siGENOME, probably the most widely used library for genome-wide RNAi screens.
siGENOME is based on low-complexity pooling, with 4 siRNAs pooled per gene. In the first pass, hits are identified based on the pooled siRNA result. In the second pass, siRNAs for hit genes are tested individually (deconvoluted).
In Falkenberg et al.’s pass 2, they examine 450 hit genes, split across 2 phenotypes of interest (Caspase-Glo 3/7 as a measure of apoptosis and Cell Fluor Titre, CTF, as a measure of general viability). They choose 317 genes for Caspase-Glo 3/7 and 150 genes for CTF (adds to 467 because 17 genes are hits for both phenotypes).
If the pooled result from pass 1 is due to an on-target effect, and if the siRNAs give consistent knockdown, the 4 siRNAs should give similar phenotypes (and they should also be similar to the pooled phenotype from pass 1).
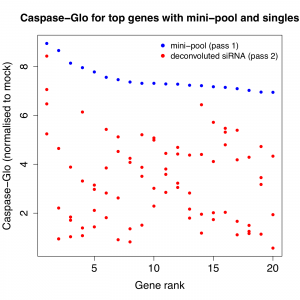
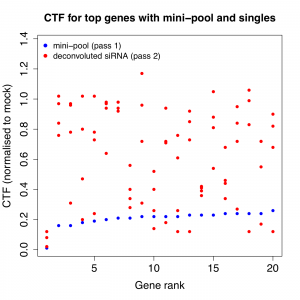
Is that what Falkenberg et al. show? Or is the pool more like Gump’s proverbial box of chocolates? The latter looks to be the case, as the following plots show. For the 2 phenotypes of interest, the top 20 genes (based on the pass 1 pool phenotype) were plotted with the pass 1 (pooled) and pass 2 (single siRNA) results.
With a few exceptions, the siRNA results are widely divergent.
(note that the pass 2 phenotypic results are weaker than pass 1, and the authors discuss the reduction in dynamic range in the second pass– however, the divergence between individual siRNAs is still pronounced)
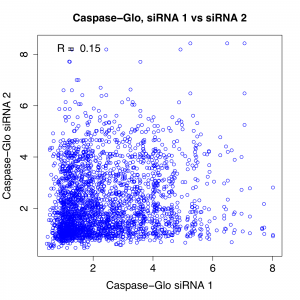
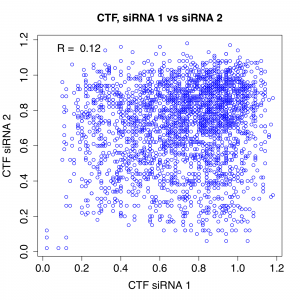
As suggested by the above plots, the correlation between different siRNAs for the same gene is very weak:
Note that R’s of 0.15 and 0.12 mean that only 2.3% and 1.4%, respectively, of the screening variance can be explained by on-target effects of the reagent.
In light of this very weak on-target signal in pass 2, it’s surprising that Dharmacon reported to the authors (after analysing their data) that there were no seed-based off-target effects. Given these results, and the known preponderance of seed-based off-target signal and the necessity of high-complexity pools to overcome seed effects, that statement must be treated very skeptically.
Overall, it looks like we can be quite confident about the top screening hit, but beyond that it really difficult to know what is causing the observed phenotype (probably not on-target knockdown effects).
To paraphrase the wisdom of Forest Gump, deconvoluted Dharmacon pools are like a box of chocolates: you never know what you’re going to get!
Additional info:
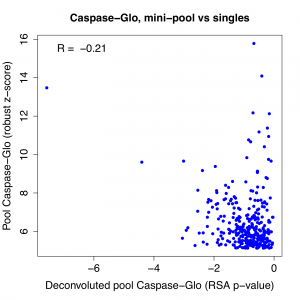
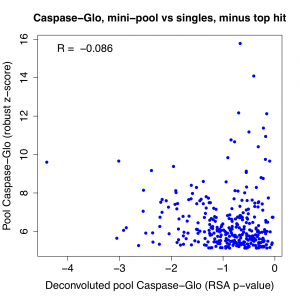
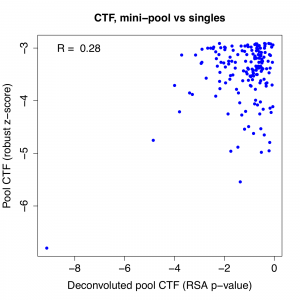
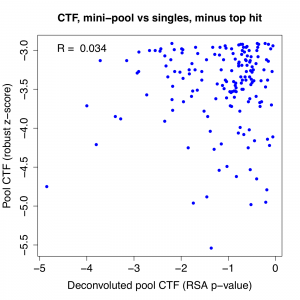
The top hit stands out when we look at the correlation between pass 1 pool result and the RSA p-value for the deconvoluted pools. We see that the correlation is quite weak to start with, and if we remove the top gene, it essentially disappears. (note: correlation is negative for Caspase-Glo because phenotypic strength is in descending direction)
Want to receive regular blog updates? Sign up for our siTOOLs Newsletter:





5 thoughts on “Deconvoluted SMARTpools are like a box of chocolates”
Comments are closed.