Pooling only 4 siRNAs increases off-target effects
In a previous post, we showed how siRNA pools with small numbers of siRNAs can exacerbate off-target effects.
Low-complexity pools (with 4 siRNAs per gene) should thus lead to overall stronger off-target effects than single siRNAs.
This phenomenon was addressed in a bioinformatics paper a few years back. The authors created a model to predict gene phenotypes based on the combined on-target and off-target effects of siRNAs.
The siRNAs were screened either individually (Ambion and Qiagen), or in pools of four (Dharmacon siGENOME), in 3 different batcterial-infection assays (B. abortus, B. henselae, and S. typhimurium).
The model assumed that each siRNA silenced its on-target gene to the same level. For off-target silencing, they used the predictions from TargetScan, a program for calculating seed-based knockdown by miRNAs or siRNAs.
In order to assess model quality, they checked how similar the gene phenotype predictions were when using different reagents types in the same pathogen-infection screen.
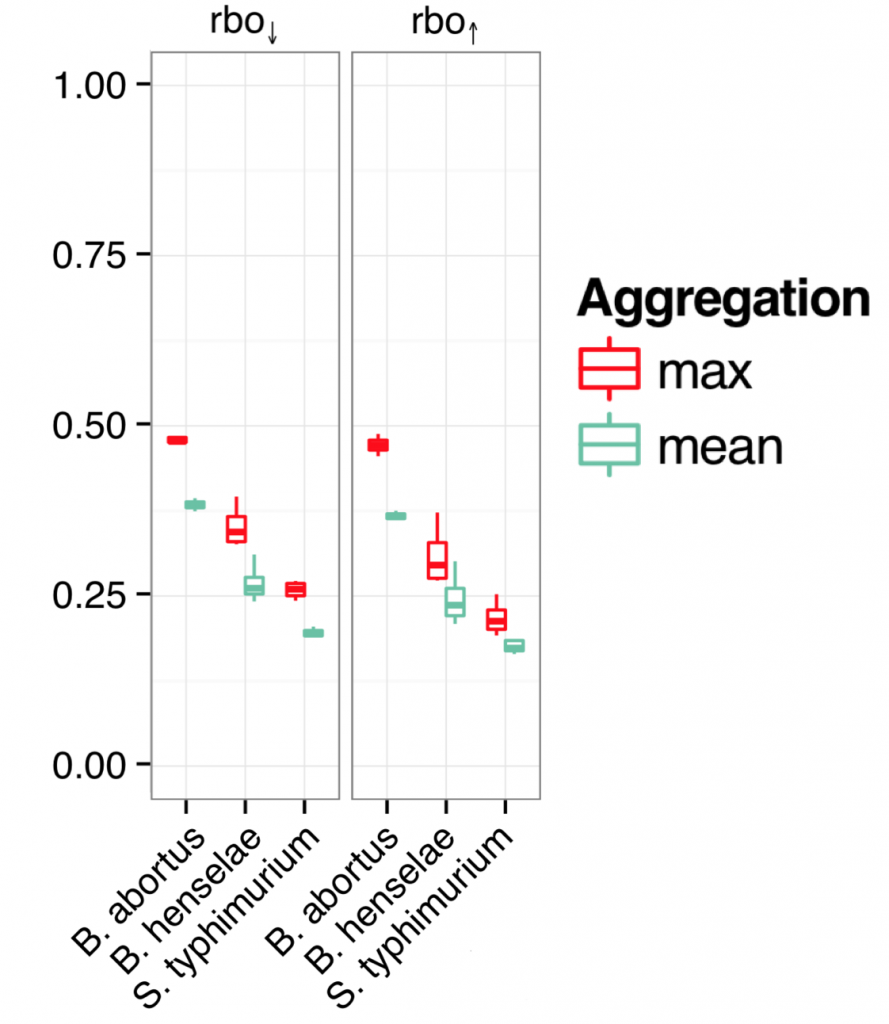
The following figure shows the rank-biased overlap (a measure of how similar lists are with regards to top- and bottom-ranked items), when estimating siGENOME off-target knockdown in one of 2 ways:
A) using the maximum TargetScan score for any of the 4 siRNAs in the siGENOME pool
B) using the mean TargetScan score for the 4 siRNAs
If low-complexity pooling increases the degree of off-target effects, we would expect the maximum TargetScan score to produce better model concordance.
And that is what the authors found. (the two plots show the rank-biased overlap for the top and bottom of the phenotype ranked lists, respectively)
The off-target effect of a 4-siRNA, low-complexity pool is best described by the strongest off-target effect of any of the individual siRNAs.
As discussed in our NAR paper, pooling a minimum of 15 siRNAs is required to reliably prevent off-target effects.